|
STA
6166 UNIT 3 Section 1 Answers
|
| Welcome | < | Begin | < | < | Unit 3 Section 1 Answers | > | Section 2 | |||||
| To Ag and Env. Answers |
| To Tox and Health Answers |
| To Social and Education Answers |
| To Engineering Answers |
Unit 3 Section 1 Answers
Toxicology and Health
From problem 9.24,page 464 in Ott and Longnecker. Researchers conducted a study of the effects of three heartworm drugs on the fat content of the shoulder muscles in Labrador retrievers. Twenty (20) dogs were randomly divided into four treatment groups of five (5) dogs each. Dogs in group A were assigned to the untreated control. The remaining three groups identify which of the heartworm drug all dogs in the group received. At the end of two years of treatment, the percentage fat content of the shoulder muscles was determined and is given here.
A B C D 2.64 3.24 2.90 2.91 2.56 3.71 3.02 2.89 3.30 2.95 3.78 3.21 2.19 3.01 2.96 2.89 2.45 3.08 2.87 2.68
- Write the linear model for these data.
- Perform a one-way analysis of variance. Test the hypothesis of no differences in average fat content at a type I error rate of 5%.
- Test the hypothesis of equal variances. Do these data need to be transformed? If so, what transformation would you use?
- For each group, compute the sample mean and subtract this mean from each observation (i.e. center each group's data). Combine all these "residuals" into on data set and test for normality. Is there any indication that these data are not normal.
- Perform the Kruskal-Wallis test on these data. How do the results compare to the analysis of variance procedure?
Answers:
1. yij = m + ai + eij
-
yij - the fat content of the j-th dog from the i-th treatment group(1=A, 2=B, 3=C, 4=D).
-
m - the overall mean fat content (if all dogs had the same treatment, its average fat content would be this.)
- ai - the (average) effect of the i-th treatment on overall average fat content- how much the i-th treatment changes the average dog's fat contentr from the overall mean, m.
- eij - the residual left over once we remove the overall mean and the effect of the i-th treatment from the (ij)-th observation - assumed to have common variance estimated by the MSE and zero mean.
2. This problem has analysis that follows closely that described in the Engineering example. Look at that problem for a detailed writeup. The SAS program for this analysis can be found here. The full output can be found here. Here I will only provide the final tables and graphs.
Dependent Variable: fat
Sum of
Source DF Squares Mean Square F Value Pr > F
Model 3 0.95052000 0.31684000 2.85 0.0703
Error 16 1.77920000 0.11120000
Corrected Total 19 2.72972000
Level of -------------fat-------------
treatment N Mean Std Dev
A 5 2.62800000 0.41227418
B 5 3.19800000 0.30605555
C 5 3.10600000 0.38115614
D 5 2.91600000 0.18942017
Bartlett's Test for Homogeneity of fat Variance
Source DF Chi-Square Pr > ChiSq
treatment 3 2.2137 0.5293
Brown and Forsythe's Test for Homogeneity of fat Variance
ANOVA of Absolute Deviations from Group Medians
Sum of Mean
Source DF Squares Square F Value Pr > F
treatment 3 0.0579 0.0193 0.26 0.8513
Error 16 1.1752 0.0734
Tests for Normality
Test --Statistic--- -----p Value-----
Shapiro-Wilk W 0.840287 Pr < W 0.004
Kolmogorov-Smirnov D 0.245416 Pr > D <0.010
Cramer-von Mises W-Sq 0.240502 Pr > W-Sq <0.005
Anderson-Darling A-Sq 1.351514 Pr > A-Sq <0.005
Kruskal-Wallis one way analysis of variance
for 2-year dog fat data for Problem 9.24
The NPAR1WAY Procedure
Wilcoxon Scores (Rank Sums) for Variable fat
Classified by Variable treatment
Sum of Expected Std Dev Mean
treatment N Scores Under H0 Under H0 Score
--------------------------------------------------------------------------
A 5 28.0 52.50 11.452131 5.60
B 5 75.0 52.50 11.452131 15.00
C 5 61.0 52.50 11.452131 12.20
D 5 46.0 52.50 11.452131 9.20
Average scores were used for ties.
Kruskal-Wallis Test
Chi-Square 6.9824
DF 3
Pr > Chi-Square 0.0725
Figure 1. Histogram of residuals with overlay of normal density curve.
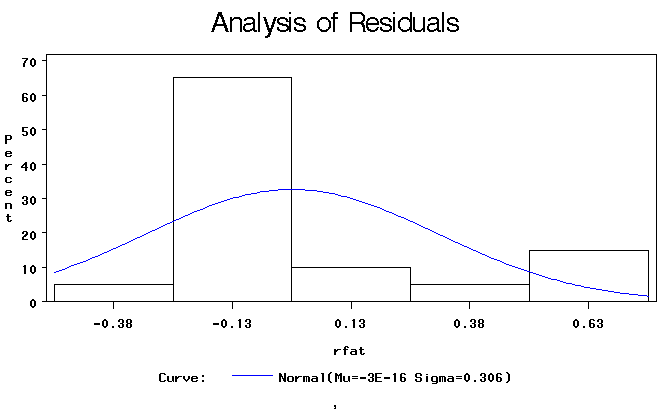
Figure 2. Normal probability plot of residuals.
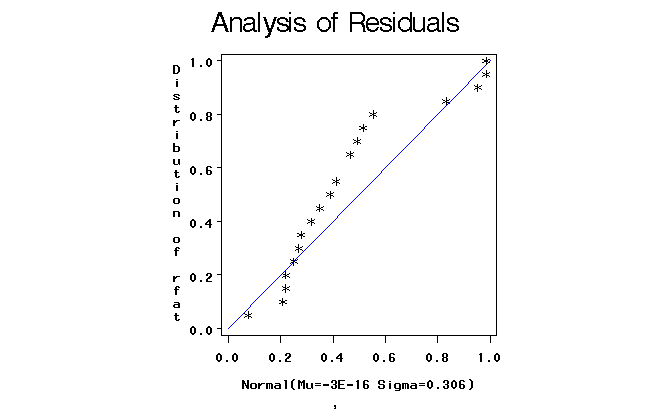
The formal normality test and the probability plot suggest that possibly these data could benifit from a transformation. But what transformation? Plotting the treatment sample means versus the sample variances or even standard deviations suggests that none of the standard transformation would work.
Figure 3. Plot of treatment sample means versus variances.
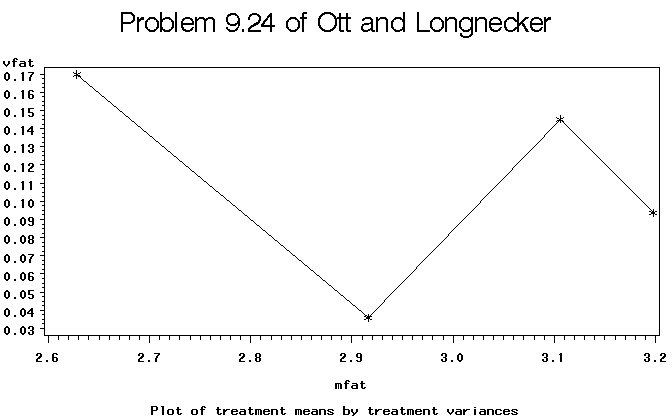
Since the tests of variances do not reject the null hypothesis of equal variances and there seems no obvious way to transform the data that would simultaneously maintain homogeneity while providing for more normally distributed residuals, we will tentatively accept the analysis of variance on the untransformed observations and conclude that there are no treatment differences (p-value = 0.07 which is greater than our a=0.05 typical test level.)
We compute the non-parameteric Kruskal-Wallis test statistics as a further check on our decision. The results of the test suggest further that there are no statistically significant differences among the treatments (p-value = 0.0725).